Category Archives: science of life
Structure-based drug design updates for SARS-CoV-2
I will be posting here information that is published regarding available crystal structures from SARS-CoV-2 and any other information relevant to structure-based drug design against the virus. Key proteins and their roles in viral infection can be found here. I hope you find this informative and useful for your research.
25 March 2020: 68 crystal structures of SARS-CoV-2 protease, with various bound fragments identified using PanDDA analysis are listed here by EBI.
25 March 2020: The Diamond Light Source (UK) has been able to solve a new structure of the SARS-CoV-2 main protease (MPro) at high resolution (1.25 Å, PDB ID: 6YB7), and subsequently complete a large XChem crystallographic fragment screen against it (detailed here). Data have been deposited with the PDB, and are made available immediately to the world on this page; additional work is ongoing, and updates will be continually posted in their website. On Tuesday March 17th they publicly released results from the full 1500-crystal experiment that yielded 58 non-covalent and covalent active-site fragments. Following data reprocessing and further analysis, an additional 13 structures were released on March 24th taking the final total to 66 active site fragments, 44 of which were covalently bound (full timeline here, download page here). This was an exceptionally large screen, and yielded an exceptionally rich readout, with vast opportunities for fragment growing and merging.
25 March 2020: Blog post, with code by Patrick Walters in Practical Cheminformatics “Building on the Fragments From the Diamond/XChem SARS-CoV-2 Main Protease (MPro) Fragment Screen (Part I)” . Share schemoinformatics techniques and the code used when working with the results of fragment screens.
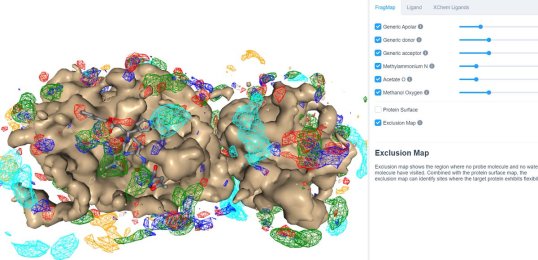
25 March 2020: Visualize SILCS FragMaps for the SARS-CoV-2 main protease (PDB ID 6LU7) through the SILCS demo web viewer, and check out the 6LU7 crystallographic ligand and the XChem fragment screen results from the Diamond Light Source.
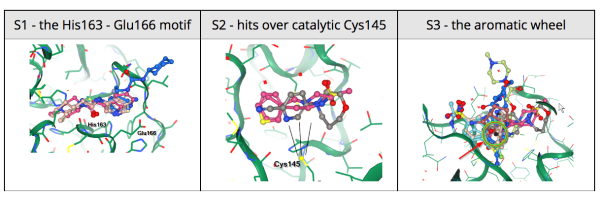
24 March 2020: Results of Diamond Light Source fragment screen:There were 68 hits of high interest – data and extensive details are here, and some interactive views here:
- 22 non-covalent hits in the active site
- 44 covalent hits in the active site
- 2 hits in the dimer interface, one in a calculated hotspot
Based on these data, the team invites chemists around the world to design new compounds or present existing compounds that could bind to the protease and submit them here.Structures submitted are prioritized by factors such as ease of synthesis and potential toxicity and the compounds selected will be synthesized and evaluated for binding to the SARS-CoV-2 protease.
 Results from the XChem fragment screen of the Diamond Light Source can be found here.
Results from the XChem fragment screen of the Diamond Light Source can be found here.
20 March 2020: The crystal structure of SARS-CoV-2 main protease is published complexed with an α-ketoamide inhibitor (PDB ID: 6Y7M). The main viral protease (Mpro, also called 3CLpro) is one of the best characterized drug targets among coronaviruses. 3CLpro protease is required for the virus but lacks human homologous proteins and thus inhibitors of this protease are less likely to bind to a human protease. Research teams in Germany have been able to crystallize the protease and have used this structure to optimize an existing α-ketoamide inhibitor developed to combat other diseases.
 SARS-CoV-2 coronavirus protease dimer bound to an α-ketoamide inhibitor (yellow). The image is reproduced from C&EN news.
SARS-CoV-2 coronavirus protease dimer bound to an α-ketoamide inhibitor (yellow). The image is reproduced from C&EN news.
20 March 2020: SwissProt has modeled the full SARS-CoV-2 proteome based on the NCBI reference sequence NC_045512 which is identical to GenBank entry MN908947, and annotations from UniProt. All data is deposited here.
20 March 2020: Check out refined coordinates of existing experimental SARS-CoV-2 structures using ISOLDE by Tristan Croll (Cambridge University). Modeled coordinates are deposited here.
20 March 2020: Prediction of 10 models for SARS-CoV-2 proteins from the Feig lab.
17 March 2020: The first vaccine clinical trial against SARS-CoV-2 starts in Seattle by Moderna in collaboration with NIAID. The experimental vaccine contains synthetic m-RNA clones that encode glycoprotein S. Once human cells recognize the genetic material of the virus, it is hoped that they will produce antibodies that inactivate the viral glycoprotein S. This strategy is different from the Influenza vaccine, where the viral surface protein itself, hemagglutinin, is introduced into the body and the human immune system produces antibodies against that protein.
5 March 2020: Check out computational predictions of protein structures associated with COVID-19 by DeepMind using AlphaFold, their recently published deep learning system, focuses on predicting protein structure accurately when no structures of similar proteins are available, called “free modelling”.
3 March 2020: Researchers at the Structural Genomic Infectious Diseases Center resolve the crystal structure of SARS-CoV-2 Nsp15 / NendoU endoribonuclease in high resolution. (PDB IDs: 6W01, 6VWW).
20 February 2020: A third research team also published the structure of the SARS-CoV-2 glycoprotein S of SARS-CoV-2 in two conformations using cryo-electron microscopy. (PDB IDs:6VXX, 6VYB)
19 February 2020: A research team from China presents cryo-EM structures of the SARS-CoV-2 glycoprotein S bound to the human ACE2 receptor. The structure shows the full-length human ACE2, in the presence of a neutral amino acid transporter B0AT1, with or without the receptor binding domain (RBD) of the surface spike glycoprotein (S protein) of SARS-CoV-2, both at an overall resolution of 2.9 Å, with a local resolution of 3.5 Å at the ACE2-RBD interface (PDB IDs: 6M18, 6M1D, 6M17).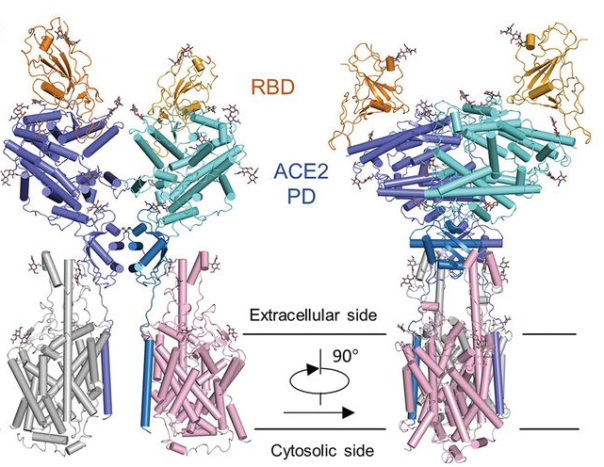 Overall structure of the RBD-ACE2-B0AT1 complex. The complex is colored by subunit, with the ACE2 protease (PD) region depicted in cyan and the Collectrin-like region (CLD) of ACE2 in blue. The glycosylated parts of the human ACE2 receptor are illustrated in strick representation. The RBD subunit of the structure of glycoprotein S that binds to ACE2 is depicted in yellow. The figure has been adapted from the original publication.
Overall structure of the RBD-ACE2-B0AT1 complex. The complex is colored by subunit, with the ACE2 protease (PD) region depicted in cyan and the Collectrin-like region (CLD) of ACE2 in blue. The glycosylated parts of the human ACE2 receptor are illustrated in strick representation. The RBD subunit of the structure of glycoprotein S that binds to ACE2 is depicted in yellow. The figure has been adapted from the original publication.
15 February 2020: The first cryo-EM structure of the viral spike protein (glycoprotein S) is published at a resolution of 3.5 Å (PDB code: 6VSB). Based on the SARS-CoV-2 genome sequence shared by Chinese researchers, they managed to prepare a purified sample of the spike protein and to determine its structure using single particle cryo-EM in less than two weeks. Using surface plasmon resonance they also measured that SARS-CoV-2 glycoprotein S binds 10-20 times more to the human cell ACE2 receptor than the 2002 SARS-CoV virus glycoprotein S. 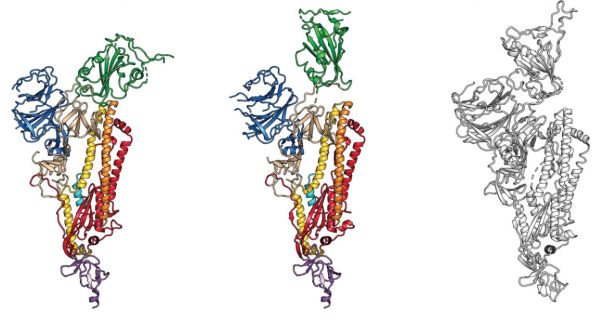 The structure of the glycoprotein S of SARS-CoV-2, as identified by cryo-electron microscopy. Green binds to the binding site that binds to human cells. In the middle image, the green binding domain of glycoprotein S is positioned for binding to the human enzyme ACE2. On the right is the SARS-CoV glycoprotein S for comparison. The figure has been adapted from the original publication.
The structure of the glycoprotein S of SARS-CoV-2, as identified by cryo-electron microscopy. Green binds to the binding site that binds to human cells. In the middle image, the green binding domain of glycoprotein S is positioned for binding to the human enzyme ACE2. On the right is the SARS-CoV glycoprotein S for comparison. The figure has been adapted from the original publication.
5 February 2020: The crystal structure of the SARS-CoV-2 main protease is released in the PDB (PDB ID: 6LU7).
3 February 2020: The complete viral genome of the new coronavirus was analyzed by scientists in China.
30 January 2020: The World Health Organization has named the new virus SARS-CoV-2 and the disease caused by the new virus, COVID-19, and the outbreak was declared a Public Health Emergency of International Concern.
5 January 2020: The source of this infection was quickly identified as a new coronavirus that resembles those that caused SARS-CoV outbreaks in 2002-2004 and MERS-CoV in 2012, but is not the same virus.
31 December 2019: A pneumonia of unknown cause detected in Wuhan, China was first reported to the WHO Country Office in China on 31 December 2019.
Pheromones are in the Air: Friends, Lovers or Foes
Love at first sight? Desire in a bottle? There have been a lot of claims about what pheromones can do, but what is the science? Dr. George Preti and Dr. Charles J. Wysocki of the Monell Chemical Senses Center discuss the chemistry of pheromones and how mood and emotion can be communicated through chemical signals.
The Chemistry of Love
I was sitting in the lobby of my accountant’s office, flipping absentmindedly through a magazine when she walked in. I’ve never had a visceral reaction as when I saw her walk through that door. There was just something about her; I felt head over heels… My heart started racing and I had butterflies in my stomach…
 This is the amazing time when you are truly love-struck. With an irresistible cocktail of chemicals, our brain entices us to fall in love. But is it really us or is it yet another nature’s trick to keep our species alive?
This is the amazing time when you are truly love-struck. With an irresistible cocktail of chemicals, our brain entices us to fall in love. But is it really us or is it yet another nature’s trick to keep our species alive?
Scientists agree that there are three stages and processes in love:
Stage 1 – Attraction : Dopamine and Adrenaline
: Dopamine and Adrenaline
When you fall in love, your brain starts sending signals before you can even blink. Your heart races and palms sweat: adrenaline is getting released from neurons. Then, when you are close to your sweetheart, dopamine is released, which triggers euphoria and feelings of bliss, increased energy, increased energy, less need for sleep or food, and focused attention on your new relationship. Phenylethylamine (PEA) is the chemical responsible for releasing adrenaline and dopamine. Biologically speaking, with the combination of dopamine, adrenaline, and PEA in your body, you’re experiencing something similar to a cocaine high. And it doesn’t matter whether you just met this *special* person or you ‘ve known them for ages; these love signals can be released at any time. Actually, no, not at any time. Only when mother nature thinks the timing is right** (see below).
Stage 2 – Lust: Testosterone kicks in
That is about the time that androgens and estrogens kick in. Estrogens and androgens are steroids, which are the female and male sex hormones, respectively. Testosterone is the sex hormone in both male and females, which is what gives us libido and sex drive. This hormone is elevated when we are in love. This makes sense as you’re most concerned about sex when it’s most likely to result in reproduction. Some people argue that this Stage precedes Stage 1, but it’s up to you to decide 🙂
Stage 3 – Attachment: Oxytocin, the love hormone
In the long run, two other hormones make their headway when emotionally involved: the “hormones of attachment”, oxytocin and vasopressin.
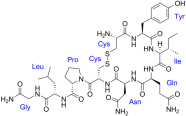
Oxytocin. Notice something? As we go down the love chain, the molecules appear more complex; sort of like love-life!
Oxytocin is a hormone stored in the brain. When the hormone is released in the brain, it results in feelings of attachment and bonding. It has been found that oxytocin gets released at orgasm, which is why couples feel much closer to one another after they have had sex. Oxytocin is also released at childbirth and is partly responsible for the strong bond between mum and baby (actually oxytocin has a Greek root: “οξυτοκίνη”, which means “quick birth”). So, if you feel the urge for a stay-at-home movie night to cuddle with your beau, it may be oxytocin at work.
Vasopressin is another important hormone when it comes to relationships. Scientists studied the prairie vole, which is known to be monogamous and to express vasopressin like humans. When the researchers introduced vasopressin receptors to the “meadow vole”, the promiscuous cousin of the prairie vole, these playboys reformed their ways and fixated on one female, choosing to mate with only her – even when other females tried to tempt them!!
Wondering why some people find it hard to get emotionally involved or committed?
Some people might be less prone to form stable relationships because they might be not express as many “oxytocin or vasopressin receptors” (proteins that take up oxytocin or vasopressin) in their brains. Thus, some people are not that sensitive to the “commitment hormones” oxytocin and vasopressin. Experiments in animals showed that if you remove the oxytocin or vasopressin receptors, these animals cannot form strong bonds with each other any more.
Timing is important. The perfect partner can sit right next to you at a party, and you might not notice him or her if you’re too busy at work, enmeshed in another relationship, or otherwise preoccupied. But if you’ve just moved to a new city, recovered from an unsatisfying love affair, begun to make enough money to raise a family, are suffering through a difficult experience, or have a good deal of spare time, you are ripe to fall in love.
So love appears to all be yet another one of nature’s traits:
Romantic love evolved to enable you to focus your energy on just one person at a time, conserving time and energy. The sex drive evolved to get you out there looking for partners. Attachment, the feeling of security you can feel with a long-term partner, evolved to help you stay together long enough to raise kids.
What are we really made of?
Watch this!! – I just found this amazing video that offers a very entertaining introduction into the particles and forces that make up me, you, the earth, the universe. Featuring Morgan Freeman, Stephen Hawking, Michio Kaku, Brian Cox, Richard Feynman, and Frank Close, this video offers a cursory introduction to quantum chemistry – the science that governs the interactions of subatomic particles, atoms, and even molecules.
Basically, in 3 and a half minutes, the video tries to explain that:
Dig deep inside the atom
and you’ll find tiny particles
Held together by invisible forces
The atoms that we’re made of have
Negatively charged electrons
Whirling around a big bulky nucleus
The universe is made of
Twelve particles of matter
Four forces of nature
The world is a dynamic mess
Of jiggling things
In the quantum world electrons behave as waves and as particles
For the ultimate triumph of science
We need a theory of everything
Which is still just beyond our grasp
The theory of everything is the Holy Grail of Physics: A theory that will explain through a single model everything from Creation, to supernovas, to atoms and molecules, perhaps even DNA, people, and love. (The four yet unified forces of nature are: gravitation, electromagnetism and the weak and strong nuclear forces that keep together elementary particles.)
If we ever create this theory, we might answer questions such as:
- What happened before the big bang?
- Is it possible to build a time machine?
- Can we punch a hole in space?
“The Quantum World” music video is the eleventh installment in the ongoing Symphony of Science music video series. Materials used in the creation of this video are from:
http://symphonyofscience.com for downloads & more videos!